Using Spatial Bayesian methods in `cityClimateHealth`
bayesian_demo.RmdAnother way that this model can be solved is by using Bayesian inference, implemented in STAN. We are including this implementation here so that it makes sense why we are including it later on.
The innovation here is combining the method of Armstrong 2014 with a spatial method, in this case BYM2
We implemented the spatial bayesian method of BYM2 but instead of regular poisson as a conditional poisson (i.e., multinomial) which has performance gains that they articulate in Amrstrong.
This requires bringing in a shapefile, so you can define the network
The standard application is using MCMC, we also include all STAN model types:
- MCMC
- laplace
- variational
- pathfinder
You can also experiment with speeding things up (at the risk of less precise estimates) using the laplace or variational method. see Jack’s notes as so what is going on here
library(data.table)
data("ma_exposure")
data("ma_deaths")
# create exposure matrix
exposure_columns <- list(
"date" = "date",
"exposure" = "tmax_C",
"geo_unit" = "TOWN20",
"geo_unit_grp" = "COUNTY20"
)
TOWNLIST <- c('CHELSEA', 'EVERETT', 'REVERE', 'MALDEN')
exposure <- subset(ma_exposure, TOWN20 %in% TOWNLIST & year(date) %in% 2012:2015)
exposure_mat <- make_exposure_matrix(exposure, exposure_columns)
#> Warning in make_exposure_matrix(exposure, exposure_columns): check about any NA, some corrections for this later,
#> but only in certain columns
# create outcome table
outcome_columns <- list(
"date" = "date",
"outcome" = "daily_deaths",
"factor" = 'age_grp',
"factor" = 'sex',
"geo_unit" = "TOWN20",
"geo_unit_grp" = "COUNTY20"
)
deaths <- subset(ma_deaths, TOWN20 %in% TOWNLIST & year(date) %in% 2012:2015)
deaths_tbl <- make_outcome_table(deaths, outcome_columns)
# plot
data("ma_towns")
library(ggplot2)
local_shp <- subset(ma_towns, TOWN20 %in% TOWNLIST)
ggplot(local_shp) + geom_sf(aes(fill = TOWN20))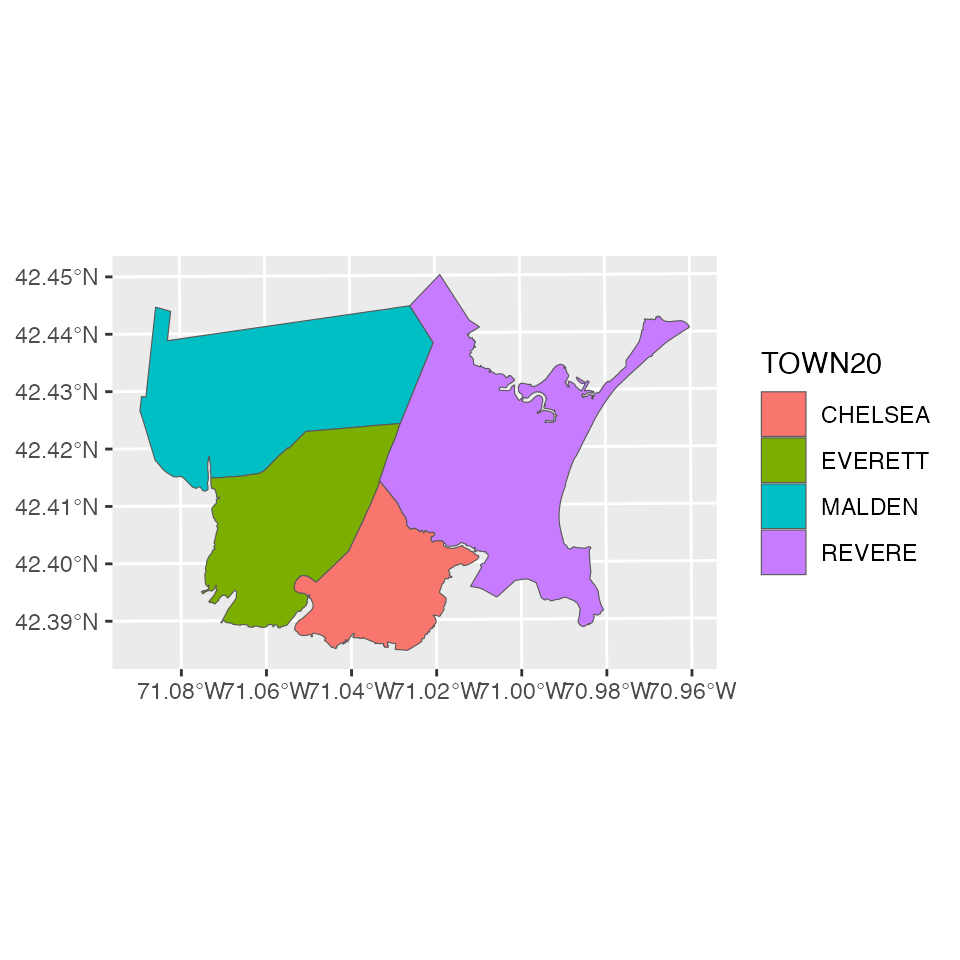
Now get initial estimates for each geo_unit
beta_l <- vector("list", 4)
cr_l <- vector("list", 4)
plot_l <- vector("list", 4)
cb_list <- vector("list", 4)
oo_list <- vector("list", 4)
for(bb in 1:4) {
m1 <- condPois_1stage(
subset(exposure_mat, TOWN20 == TOWNLIST[bb]),
subset(deaths_tbl, TOWN20 == TOWNLIST[bb]),
global_cen = 15)
cb_list[[bb]] <- m1$`_`$out[[1]]$orig_basis
oo_list[[bb]] <- m1$`_`$out[[1]]$outcomes
beta_l[[bb]] <- m1$`_`$out[[1]]$orig_coef
cr_l[[bb]] <- m1$`_`$out[[1]]$coef
plot_l[[bb]] <- plot(m1)
}
#>
#> crossbasis args:
#>
#> maxlag: 5
#>
#> argvar:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: Named num [1:2] 25.8 31.3
#> ..- attr(*, "names")= chr [1:2] "50%" "90%"
#>
#> arglag:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: num [1:2] 0.878 2.095
#>
#> strata:
#> CHELSEA:yr2012:mn05:dow03
#> strata_min: 0
#>
#>
#> crossbasis args:
#>
#> maxlag: 5
#>
#> argvar:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: Named num [1:2] 25.8 31
#> ..- attr(*, "names")= chr [1:2] "50%" "90%"
#>
#> arglag:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: num [1:2] 0.878 2.095
#>
#> strata:
#> EVERETT:yr2012:mn05:dow03
#> strata_min: 0
#>
#>
#> crossbasis args:
#>
#> maxlag: 5
#>
#> argvar:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: Named num [1:2] 25.1 30.1
#> ..- attr(*, "names")= chr [1:2] "50%" "90%"
#>
#> arglag:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: num [1:2] 0.878 2.095
#>
#> strata:
#> REVERE:yr2012:mn05:dow03
#> strata_min: 0
#>
#>
#> crossbasis args:
#>
#> maxlag: 5
#>
#> argvar:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: Named num [1:2] 24.4 29
#> ..- attr(*, "names")= chr [1:2] "50%" "90%"
#>
#> arglag:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: num [1:2] 0.878 2.095
#>
#> strata:
#> MALDEN:yr2012:mn05:dow03
#> strata_min: 0
mx <- do.call(cbind, beta_l) # COEFS NOT THE SAME
colnames(mx) = TOWNLIST
mx
#> CHELSEA EVERETT REVERE MALDEN
#> cbv1.l1 0.0451309468 0.05298088 -0.0005224117 0.017711625
#> cbv1.l2 -0.0223211761 -0.03592669 0.0184129771 0.012490790
#> cbv1.l3 0.0873424413 0.08593669 0.1029649763 0.093454486
#> cbv1.l4 -0.0335859603 -0.03578575 -0.0337628164 -0.041517343
#> cbv2.l1 0.0852368471 0.05049362 -0.0533811289 0.039648824
#> cbv2.l2 0.0166508607 -0.11934992 0.0747314454 0.055829432
#> cbv2.l3 0.1926712791 0.18510476 0.1393985118 0.118837565
#> cbv2.l4 -0.0462482063 -0.04542133 -0.0608086198 -0.054356689
#> cbv3.l1 0.0170645385 0.07301390 0.0257165131 0.038563344
#> cbv3.l2 -0.0005616437 -0.06948122 0.0223225434 -0.002010598
#> cbv3.l3 0.1718665757 0.12512136 0.1186064940 0.130222731
#> cbv3.l4 -0.0490847643 -0.01913613 -0.0655887549 -0.044893272
mcr <- do.call(cbind, cr_l) # COEFS THE SAME
colnames(mcr) = TOWNLIST
mcr
#> CHELSEA EVERETT REVERE MALDEN
#> b1 0.1847675 0.1774954 0.2034421 0.1922760
#> b2 0.4637942 0.3065562 0.2523009 0.2921484
#> b3 0.3375227 0.2624969 0.2438566 0.2749740
library(patchwork)
wrap_plots(plot_l)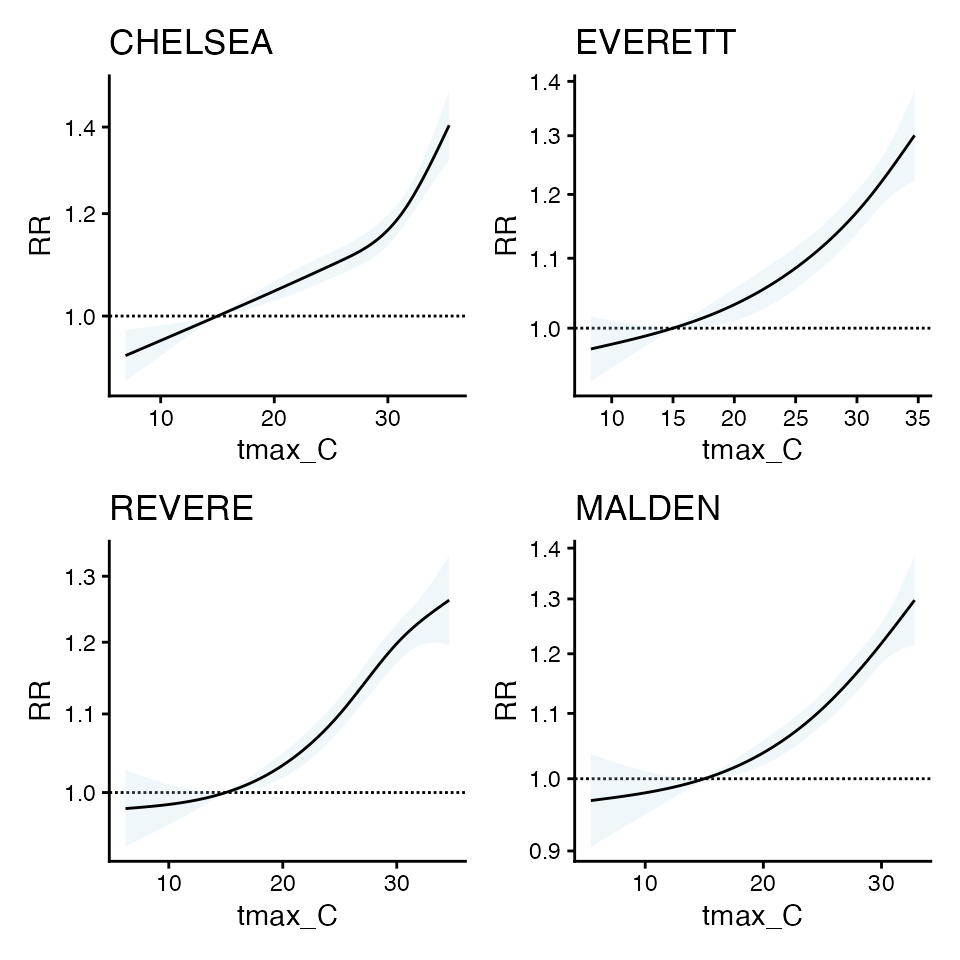
the cr coefs are similar
the orig_coefs are not, which is why beta-wise implementation of SB_DLNM method doesn’t work - because the don’t have to be the same to produce similar curves.
So, instead of forcing Beta to be similar, we can use bym2
refs:
- https://mc-stan.org/learn-stan/case-studies/icar_stan.html
- https://link.springer.com/article/10.1186/1476-072X-4-31
- https://github.com/stan-dev/example-models/blob/e5b7d9e2e9ecc375805c7e49e4a4d4c1882b5e3b/knitr/car-iar-poisson/bym2_predictor_plus_offset.stan#L4
ok here’s the ref of how LAPLACE works:
- https://mc-stan.org/cmdstanr/reference/model-method-laplace.html
- https://statmodeling.stat.columbia.edu/2023/02/08/implementing-laplace-approximation-in-stan-whats-happening-under-the-hood/
I think this makes for a good candidate because betas are normal and the model is not hierarchical
m_sb1 <- condPois_sb(exposure_mat, deaths_tbl, local_shp,
stan_type = 'mcmc',
verbose = 2,
global_cen = 15,
stan_opts = list(refresh = 200),
use_spatial_model = 'none')
#> STAN TYPE = mcmc
#> SPATIAL MODEL = none
#> -- validation passed
#> -- prepare inputs
#> CHELSEA
#> crossbasis args:
#>
#> maxlag: 5
#>
#> argvar:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: Named num [1:2] 25.8 31.3
#> ..- attr(*, "names")= chr [1:2] "50%" "90%"
#>
#> arglag:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: num [1:2] 0.878 2.095
#>
#> strata:
#> CHELSEA:yr2012:mn05:dow03
#> strata_min: 0
#> Warning in condPois_1stage(exposure_matrix = single_exposure_matrix,
#> outcomes_tbl = single_outcomes_tbl, : Centering point is outside the range of
#> exposures in geo-unit CHELSEA. This means your zones are across too large of an
#> area, or there are differences in exposures so much that the bases are quite
#> different. Try limiting the geo-units passed in to those that are more similar,
#> manually setting a centering point that you know each geo-unit has, or changing
#> your exposure variable.
#> EVERETT MALDEN REVERE
#> Warning in condPois_1stage(exposure_matrix = single_exposure_matrix,
#> outcomes_tbl = single_outcomes_tbl, : Centering point is outside the range of
#> exposures in geo-unit REVERE. This means your zones are across too large of an
#> area, or there are differences in exposures so much that the bases are quite
#> different. Try limiting the geo-units passed in to those that are more similar,
#> manually setting a centering point that you know each geo-unit has, or changing
#> your exposure variable.
#> Warning in getSW(shp = shp_sf_safe, ni = 1, include_self = F): has to be one
#> polygon per row in `shp`
#>
#> -- run STAN
#> ...mcmc...
#> Running MCMC with 2 parallel chains...
#>
#> Chain 1 Iteration: 1 / 2000 [ 0%] (Warmup)
#> Chain 2 Iteration: 1 / 2000 [ 0%] (Warmup)
#> Chain 2 Iteration: 200 / 2000 [ 10%] (Warmup)
#> Chain 1 Iteration: 200 / 2000 [ 10%] (Warmup)
#> Chain 2 Iteration: 400 / 2000 [ 20%] (Warmup)
#> Chain 1 Iteration: 400 / 2000 [ 20%] (Warmup)
#> Chain 2 Iteration: 600 / 2000 [ 30%] (Warmup)
#> Chain 1 Iteration: 600 / 2000 [ 30%] (Warmup)
#> Chain 2 Iteration: 800 / 2000 [ 40%] (Warmup)
#> Chain 1 Iteration: 800 / 2000 [ 40%] (Warmup)
#> Chain 2 Iteration: 1000 / 2000 [ 50%] (Warmup)
#> Chain 2 Iteration: 1001 / 2000 [ 50%] (Sampling)
#> Chain 1 Iteration: 1000 / 2000 [ 50%] (Warmup)
#> Chain 1 Iteration: 1001 / 2000 [ 50%] (Sampling)
#> Chain 2 Iteration: 1200 / 2000 [ 60%] (Sampling)
#> Chain 1 Iteration: 1200 / 2000 [ 60%] (Sampling)
#> Chain 2 Iteration: 1400 / 2000 [ 70%] (Sampling)
#> Chain 1 Iteration: 1400 / 2000 [ 70%] (Sampling)
#> Chain 2 Iteration: 1600 / 2000 [ 80%] (Sampling)
#> Chain 1 Iteration: 1600 / 2000 [ 80%] (Sampling)
#> Chain 2 Iteration: 1800 / 2000 [ 90%] (Sampling)
#> Chain 1 Iteration: 1800 / 2000 [ 90%] (Sampling)
#> Chain 2 Iteration: 2000 / 2000 [100%] (Sampling)
#> Chain 2 finished in 28.8 seconds.
#> Chain 1 Iteration: 2000 / 2000 [100%] (Sampling)
#> Chain 1 finished in 29.0 seconds.
#>
#> Both chains finished successfully.
#> Mean chain execution time: 28.9 seconds.
#> Total execution time: 29.1 seconds.
#>
#> ...mcmc draws...
#> CHELSEA EVERETT MALDEN REVERE
#> -- apply estimatesCompare, first you can see that with spatial_model = F, there is similarity in beta coefs
mx
#> CHELSEA EVERETT REVERE MALDEN
#> cbv1.l1 0.0451309468 0.05298088 -0.0005224117 0.017711625
#> cbv1.l2 -0.0223211761 -0.03592669 0.0184129771 0.012490790
#> cbv1.l3 0.0873424413 0.08593669 0.1029649763 0.093454486
#> cbv1.l4 -0.0335859603 -0.03578575 -0.0337628164 -0.041517343
#> cbv2.l1 0.0852368471 0.05049362 -0.0533811289 0.039648824
#> cbv2.l2 0.0166508607 -0.11934992 0.0747314454 0.055829432
#> cbv2.l3 0.1926712791 0.18510476 0.1393985118 0.118837565
#> cbv2.l4 -0.0462482063 -0.04542133 -0.0608086198 -0.054356689
#> cbv3.l1 0.0170645385 0.07301390 0.0257165131 0.038563344
#> cbv3.l2 -0.0005616437 -0.06948122 0.0223225434 -0.002010598
#> cbv3.l3 0.1718665757 0.12512136 0.1186064940 0.130222731
#> cbv3.l4 -0.0490847643 -0.01913613 -0.0655887549 -0.044893272
m_sb1$`_`$beta_mat
#> CHELSEA EVERETT MALDEN REVERE
#> [1,] 0.04563081 0.05352101 0.01681938 -0.0007161936
#> [2,] -0.02310974 -0.03545623 0.01349965 0.0180104763
#> [3,] 0.08724563 0.08576537 0.09334545 0.1028893168
#> [4,] -0.03350999 -0.03532395 -0.04105760 -0.0331877843
#> [5,] 0.08698833 0.04810957 0.03598086 -0.0511078793
#> [6,] 0.01534864 -0.11608512 0.06032731 0.0744266904
#> [7,] 0.19255327 0.18545340 0.11679099 0.1364912451
#> [8,] -0.04584014 -0.04524805 -0.05158503 -0.0596107002
#> [9,] 0.01789267 0.07015688 0.03754349 0.0284379689
#> [10,] -0.00093469 -0.06706039 -0.00138018 0.0211865836
#> [11,] 0.17143376 0.12552996 0.12973764 0.1171559579
#> [12,] -0.04891770 -0.02000770 -0.04463672 -0.0651447948Compare with spatial model
using laplace in this case, but you could try mcmc
m_sb2 <- condPois_sb(exposure_mat, deaths_tbl, local_shp,
stan_type = 'laplace',
verbose = 2,
global_cen = 15,
stan_opts = list(refresh = 200),
use_spatial_model = 'bym2')
#> STAN TYPE = laplace
#> SPATIAL MODEL = bym2
#> -- validation passed
#> -- prepare inputs
#> CHELSEA
#> crossbasis args:
#>
#> maxlag: 5
#>
#> argvar:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: Named num [1:2] 25.8 31.3
#> ..- attr(*, "names")= chr [1:2] "50%" "90%"
#>
#> arglag:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: num [1:2] 0.878 2.095
#>
#> strata:
#> CHELSEA:yr2012:mn05:dow03
#> strata_min: 0
#> Warning in condPois_1stage(exposure_matrix = single_exposure_matrix,
#> outcomes_tbl = single_outcomes_tbl, : Centering point is outside the range of
#> exposures in geo-unit CHELSEA. This means your zones are across too large of an
#> area, or there are differences in exposures so much that the bases are quite
#> different. Try limiting the geo-units passed in to those that are more similar,
#> manually setting a centering point that you know each geo-unit has, or changing
#> your exposure variable.
#> EVERETT MALDEN REVERE
#> Warning in condPois_1stage(exposure_matrix = single_exposure_matrix,
#> outcomes_tbl = single_outcomes_tbl, : Centering point is outside the range of
#> exposures in geo-unit REVERE. This means your zones are across too large of an
#> area, or there are differences in exposures so much that the bases are quite
#> different. Try limiting the geo-units passed in to those that are more similar,
#> manually setting a centering point that you know each geo-unit has, or changing
#> your exposure variable.
#> Warning in getSW(shp = shp_sf_safe, ni = 1, include_self = F): has to be one
#> polygon per row in `shp`
#>
#> -- run STAN
#> ...laplace optimize...
#> Initial log joint probability = -6739.06
#> Iter log prob ||dx|| ||grad|| alpha alpha0 # evals Notes
#> 139 -6737.26 0.000348121 0.731762 1 1 169
#> Optimization terminated normally:
#> Convergence detected: relative gradient magnitude is below tolerance
#> Finished in 0.1 seconds.
#> ...laplace sample...
#> Calculating Hessian
#> Calculating inverse of Cholesky factor
#> Generating draws
#> iteration: 0
#> iteration: 100
#> iteration: 200
#> iteration: 300
#> iteration: 400
#> iteration: 500
#> iteration: 600
#> iteration: 700
#> iteration: 800
#> iteration: 900
#> Finished in 0.9 seconds.
#> ...laplace draws...
#> CHELSEA EVERETT MALDEN REVERE
#> -- apply estimatesCompare, now you can see these are different
mx
#> CHELSEA EVERETT REVERE MALDEN
#> cbv1.l1 0.0451309468 0.05298088 -0.0005224117 0.017711625
#> cbv1.l2 -0.0223211761 -0.03592669 0.0184129771 0.012490790
#> cbv1.l3 0.0873424413 0.08593669 0.1029649763 0.093454486
#> cbv1.l4 -0.0335859603 -0.03578575 -0.0337628164 -0.041517343
#> cbv2.l1 0.0852368471 0.05049362 -0.0533811289 0.039648824
#> cbv2.l2 0.0166508607 -0.11934992 0.0747314454 0.055829432
#> cbv2.l3 0.1926712791 0.18510476 0.1393985118 0.118837565
#> cbv2.l4 -0.0462482063 -0.04542133 -0.0608086198 -0.054356689
#> cbv3.l1 0.0170645385 0.07301390 0.0257165131 0.038563344
#> cbv3.l2 -0.0005616437 -0.06948122 0.0223225434 -0.002010598
#> cbv3.l3 0.1718665757 0.12512136 0.1186064940 0.130222731
#> cbv3.l4 -0.0490847643 -0.01913613 -0.0655887549 -0.044893272
m_sb2$`_`$beta_mat
#> CHELSEA EVERETT MALDEN REVERE
#> [1,] 0.045423333 0.05112039 1.900732e-02 0.0003361363
#> [2,] -0.021520151 -0.03348218 1.168377e-02 0.0171357910
#> [3,] 0.086169530 0.08576237 9.202635e-02 0.1026162647
#> [4,] -0.033034684 -0.03701113 -4.046737e-02 -0.0336513220
#> [5,] 0.088049683 0.04858261 3.603389e-02 -0.0514857925
#> [6,] 0.019462978 -0.11101076 6.234808e-02 0.0752953200
#> [7,] 0.187765616 0.18230330 1.161435e-01 0.1368418551
#> [8,] -0.042253687 -0.04710544 -5.393741e-02 -0.0612077042
#> [9,] 0.016094283 0.07222675 3.729183e-02 0.0263192109
#> [10,] 0.001261148 -0.06617505 4.976655e-05 0.0224012337
#> [11,] 0.172412859 0.12429712 1.292206e-01 0.1186910900
#> [12,] -0.048785974 -0.01986296 -4.510412e-02 -0.0658182839Compare with spatial model for leroux
using laplace in this case, but you could try mcmc
m_sb3 <- condPois_sb(exposure_mat,
deaths_tbl, local_shp,
stan_type = 'laplace',
verbose = 2,
stan_opts = list(refresh = 200),
use_spatial_model = 'leroux')
#> STAN TYPE = laplace
#> SPATIAL MODEL = leroux
#> -- validation passed
#> -- prepare inputs
#> CHELSEA
#> crossbasis args:
#>
#> maxlag: 5
#>
#> argvar:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: Named num [1:2] 25.8 31.3
#> ..- attr(*, "names")= chr [1:2] "50%" "90%"
#>
#> arglag:
#> List of 2
#> $ fun : chr "ns"
#> $ knots: num [1:2] 0.878 2.095
#>
#> strata:
#> CHELSEA:yr2012:mn05:dow03
#> strata_min: 0
#> Warning in condPois_1stage(exposure_matrix = single_exposure_matrix,
#> outcomes_tbl = single_outcomes_tbl, : Centering point is outside the range of
#> exposures in geo-unit CHELSEA. This means your zones are across too large of an
#> area, or there are differences in exposures so much that the bases are quite
#> different. Try limiting the geo-units passed in to those that are more similar,
#> manually setting a centering point that you know each geo-unit has, or changing
#> your exposure variable.
#> EVERETT MALDEN REVERE
#> Warning in condPois_1stage(exposure_matrix = single_exposure_matrix,
#> outcomes_tbl = single_outcomes_tbl, : Centering point is outside the range of
#> exposures in geo-unit REVERE. This means your zones are across too large of an
#> area, or there are differences in exposures so much that the bases are quite
#> different. Try limiting the geo-units passed in to those that are more similar,
#> manually setting a centering point that you know each geo-unit has, or changing
#> your exposure variable.
#> Warning in getSW(shp = shp_sf_safe, ni = 1, include_self = F): has to be one
#> polygon per row in `shp`
#>
#> -- run STAN
#> ...laplace optimize...
#> Initial log joint probability = -7000.25
#> Iter log prob ||dx|| ||grad|| alpha alpha0 # evals Notes
#> 199 -6518.16 0.0317223 386.68 1 1 215
#> Iter log prob ||dx|| ||grad|| alpha alpha0 # evals Notes
#> 399 -6438.84 0.0763665 11799.7 0.5406 1 425
#> Iter log prob ||dx|| ||grad|| alpha alpha0 # evals Notes
#> 599 -6393.62 0.027489 12270.4 1 1 635
#> Iter log prob ||dx|| ||grad|| alpha alpha0 # evals Notes
#> 799 -6371.4 0.00210131 22054.6 1 1 849
#> Iter log prob ||dx|| ||grad|| alpha alpha0 # evals Notes
#> 999 -6362.85 2.48342e-06 1876.72 1 1 1060
#> Iter log prob ||dx|| ||grad|| alpha alpha0 # evals Notes
#> 1199 -6360.11 1.70314e-06 1926.29 1 1 1272
#> Iter log prob ||dx|| ||grad|| alpha alpha0 # evals Notes
#> 1399 -6359.2 1.63738e-06 1871.19 1 1 1480
#> Iter log prob ||dx|| ||grad|| alpha alpha0 # evals Notes
#> 1599 -6358.77 1.14602e-05 1174.76 1 1 1690
#> Iter log prob ||dx|| ||grad|| alpha alpha0 # evals Notes
#> 1799 -6357.61 0.000191241 7846.74 1 1 1897
#> Iter log prob ||dx|| ||grad|| alpha alpha0 # evals Notes
#> 1999 -6356.56 1.89901e-05 1981.71 1 1 2106
#> Iter log prob ||dx|| ||grad|| alpha alpha0 # evals Notes
#> 2152 -6355.97 2.89835e-07 242.602 1 1 2271
#> Optimization terminated normally:
#> Convergence detected: relative gradient magnitude is below tolerance
#> Finished in 1.1 seconds.
#> ...laplace sample...
#> Calculating Hessian
#> Calculating inverse of Cholesky factor
#> Generating draws
#> iteration: 0
#> iteration: 100
#> iteration: 200
#> iteration: 300
#> iteration: 400
#> iteration: 500
#> iteration: 600
#> iteration: 700
#> iteration: 800
#> iteration: 900
#> Finished in 0.7 seconds.
#> ...laplace draws...
#> CHELSEA EVERETT MALDEN REVERE
#> -- apply estimatesAs you can see, lots of smoothing to a central estimate !
mx
#> CHELSEA EVERETT REVERE MALDEN
#> cbv1.l1 0.0451309468 0.05298088 -0.0005224117 0.017711625
#> cbv1.l2 -0.0223211761 -0.03592669 0.0184129771 0.012490790
#> cbv1.l3 0.0873424413 0.08593669 0.1029649763 0.093454486
#> cbv1.l4 -0.0335859603 -0.03578575 -0.0337628164 -0.041517343
#> cbv2.l1 0.0852368471 0.05049362 -0.0533811289 0.039648824
#> cbv2.l2 0.0166508607 -0.11934992 0.0747314454 0.055829432
#> cbv2.l3 0.1926712791 0.18510476 0.1393985118 0.118837565
#> cbv2.l4 -0.0462482063 -0.04542133 -0.0608086198 -0.054356689
#> cbv3.l1 0.0170645385 0.07301390 0.0257165131 0.038563344
#> cbv3.l2 -0.0005616437 -0.06948122 0.0223225434 -0.002010598
#> cbv3.l3 0.1718665757 0.12512136 0.1186064940 0.130222731
#> cbv3.l4 -0.0490847643 -0.01913613 -0.0655887549 -0.044893272
m_sb3$`_`$beta_mat
#> CHELSEA EVERETT MALDEN REVERE
#> [1,] 0.007393641 0.007393784 0.007394247 0.007393496
#> [2,] 0.005909126 0.005908665 0.005909324 0.005909028
#> [3,] 0.101804062 0.101805289 0.101804801 0.101804395
#> [4,] -0.044028808 -0.044029393 -0.044029676 -0.044029059
#> [5,] -0.045403324 -0.045399940 -0.045397528 -0.045397372
#> [6,] 0.046002454 0.046006165 0.046005329 0.046004261
#> [7,] 0.187302489 0.187302192 0.187301543 0.187302563
#> [8,] -0.079967398 -0.079968486 -0.079967632 -0.079968145
#> [9,] 0.009569402 0.009569678 0.009569970 0.009570108
#> [10,] 0.011026743 0.011027348 0.011027609 0.011027644
#> [11,] 0.141515048 0.141515237 0.141514517 0.141515253
#> [12,] -0.056311461 -0.056311481 -0.056310680 -0.056311312And you can also see that the leroux q value is quite
high
subset(m_sb3$`_`$stan_summary, variable == 'q')
#> # A tibble: 1 × 7
#> variable mean median sd mad q5 q95
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 q 0.631 0.640 0.113 0.112 0.437 0.808All of the other objects associated with condPois_1stage
or condPois_2stage will also work here, along with the
_list and factor coding